 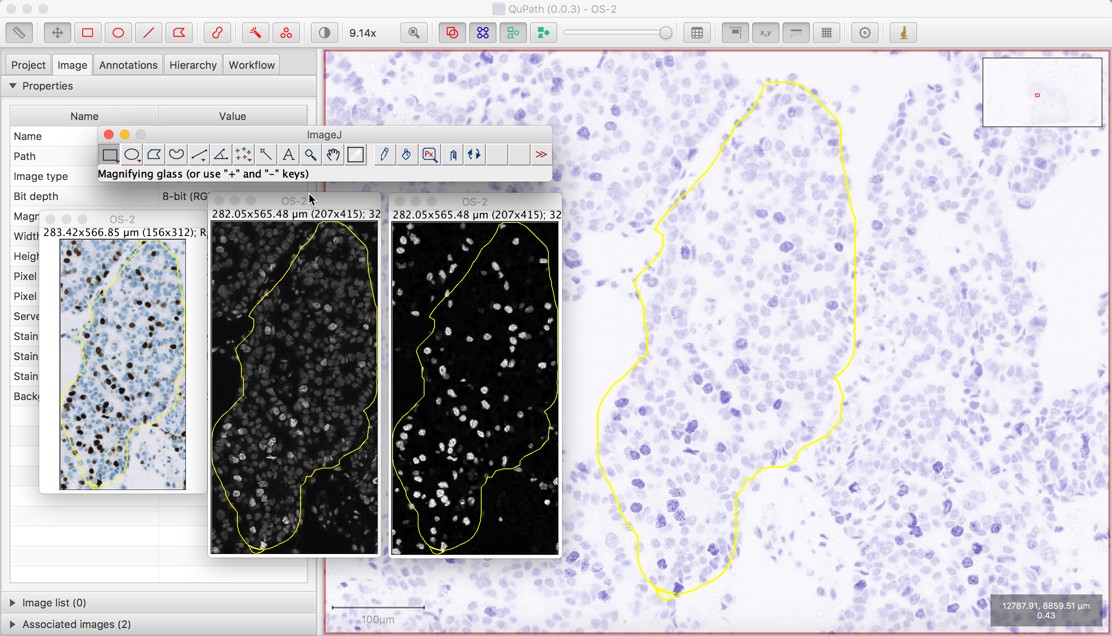
Now split out the channels into separate image-stacks.Use ImageJ/Image/Scale to resize image by either a fraction of original (0.0-1.0) or by setting the new X&Y dimensions you want (SCALE (you can see full image, but at a lower resolution).Now all channels are cropped and aligned.Use ImageJ rectangular selection tool to select out the 1000×1000 section of the image that most interests you.CROP (keeps full resolution but you can only look at a section of full image).If image more than 1000x1000x100 you may need to reduce image size so that it will load into your VR system’s GPU.Load your raw image file from your microscope into ImageJ using the Bio-Formats macro.Use ImageJ (Fiji) to create a directory of image-stack files, one file for each channel expansion microscopy), use the method below to create a directory holding one 3D image for each channel you want to view and manipulate. While single channel RGB image-stacks (z-stacks) can be loaded and viewed in ConfocalVR, if you want to manipulate the RGB colors separately, or if you are using a microscopy technique that generates multiple image stacks from the same sample (e.g. Those interested in learning more about the method can visit Wikipedia and read through the original article ( Zuiderveld, 1994).(Note – An ImageJ macro to perform this sequence is given at the end of this document) In addition, CLAHE may accentuate unwanted artifacts like beam hardening, so there is further motivation to minimize such scanning artifacts. Potential issues with CLAHE include the increase in file size associated with CT data from having both original and modified image stacks. As mentioned above, CLAHE is particularly useful for sub-optimally stained specimens, which is helpful when one cannot devote time or reserve frequent CT scanning sessions for checking and optimizing the stain concentration and duration. Generally, it improves edge recognition for all 3-D rendering programs, thus, greatly reducing the time spent on segmenting ROIs. Here is a before and after example to illustrate how CLAHE enhances diceCT images.ĭiceCT, two-year old Alligator mississippiensis head in transverse views (left) unmodified, and (right) filtered using CLAHE.ĬLAHE processes image stacks fairly quickly, so I recommend trying it with all diceCT image stacks. In CT data processing program of your choice (e.g., Avizo, VGStudio), read in the modified image stack.Once CLAHE has gone through the entire stack, save the new image stack in a different folder to keep the original CT image stack intact (you never know when you’ll need them).One can specify the “block size,” “histogram bins,” and “max slope” parameters (the details of which are outlined on ), but I have found that the default parameters do a fine job. In the text field, paste the CLAHE script. In FIJI, go to “Plugins” > “New” > “Macro”.Copy the CLAHE script from the “ Tips” section on the website.Wait until FIJI reads in the entire image stack. Drag and drop the folder that contains a stack of CT images into FIJI (download here: ).It is particularly helpful when applied to sub-optimally stained specimens.ĬLAHE is implemented in FIJI ( ImageJ) and the script is available freely and openly. This results in digital contrast enhancement that is not dominated by overly deep blacks or excessively bright whites.įor diceCT, CLAHE is very useful for improving edge recognition for digitally segmenting regions of interest (ROI) based on your CT data. In contrast to standard histogram equalization that applies single formula for enhancing contrast across the entire image, CLAHE applies multiple equalizations within partitions of an image, resulting in more localized and subtle contrast enhancements. 
By Aki Watanabe ( limited adaptive histogram equalization ( CLAHE) is a procedure for enhancing local contrast in an image or stack of images.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
